library(tidyverse)
library(janitor)
library(ggthemes)
library(rmarkdown)
library(rpart)
library(rpart.plot)
library(ranger)
library(vip)
library(pdp)
# library(xgboost) # load this right before we use xgboost()Random Forest and Gradient-Boosted Trees
Boston Housing Data
Boston Housing Data
boston <- read_csv("https://bcdanl.github.io/data/boston.csv") |>
janitor::clean_names()
paged_table(boston)Modeling goal
- Predict
medv, median housing value in thousands of dollars - Boston housing is a classic regression example with multiple socioeconomic and neighborhood predictors
- We will use it to study overfitting, pruning, random forests, and boosting
Create training and test data
set.seed(42120532)
boston <- boston |>
mutate(rnd = runif(n()))
dtrain <- boston |>
filter(rnd > .2) |>
select(-rnd)
dtest <- boston |>
filter(rnd <= .2) |>
select(-rnd)
x_train <- dtrain |> select(-medv)
y_train <- dtrain$medv
x_test <- dtest |> select(-medv)
y_test <- dtest$medvRandom Forest
Fit an initial random forest on Boston housing
set.seed(1917)
init_boston_rf <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
importance = "permutation"
)
init_boston_rfRanger result
Call:
ranger(medv ~ ., data = dtrain, num.trees = 300, importance = "permutation")
Type: Regression
Number of trees: 300
Sample size: 392
Number of independent variables: 13
Mtry: 3
Target node size: 5
Variable importance mode: permutation
Splitrule: variance
OOB prediction error (MSE): 11.83664
R squared (OOB): 0.8582419 Important ranger() arguments
init_boston_rf$num.trees[1] 300init_boston_rf$mtry[1] 3init_boston_rf$prediction.error[1] 11.83664init_boston_rf$min.node.size[1] 5init_boston_rf$importance[1] "permutation"num.trees: number of trees in the forestmtry: number of predictors considered at each splitprediction.error: out-of-bag error estimate.min.node.size: minimum terminal node sizeimportance: method used to calculate variable importance"none": do not compute variable importance (default)"impurity": compute importance based on total impurity reduction from splits"permutation": compute importance based on how much model performance worsens when a variable is shuffled
Interpreting the ranger output
Tune mtry
No loop
rf_2 <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
mtry = 2,
importance = "permutation"
)
rf_4 <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
mtry = 4,
importance = "permutation"
)
rf_6 <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
mtry = 6,
importance = "permutation"
)
rf_8 <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
mtry = 8,
importance = "permutation"
)
rf_10 <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
mtry = 10,
importance = "permutation"
)
rf_12 <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300,
mtry = 12,
importance = "permutation"
)
rf_results <- tibble(
mtry = c(2, 4, 6, 8, 10, 12),
oob_error = c(
rf_2$prediction.error,
rf_4$prediction.error,
rf_6$prediction.error,
rf_8$prediction.error,
rf_10$prediction.error,
rf_12$prediction.error
)
)
rf_results |>
paged_table()Loop
# p = the total number of predictor variables in the training data
p <- ncol(x_train)
# Create a tuning grid for mtry:
# mtry is the number of predictors randomly considered at each split
# Here, we try even-numbered values from 2 up to p
rf_grid <- tibble(mtry = seq(2, p, by = 2))
# Create an empty tibble to store the tuning results
# We will save each mtry value and its corresponding OOB error
rf_results <- tibble(
mtry = numeric(),
oob_error = numeric()
)
# Loop over each candidate mtry value
for (m in rf_grid$mtry) {
# Fit a random forest model with the current mtry value
fit <- ranger(
medv ~ .,
data = dtrain,
num.trees = 300, # grow 300 trees in the forest
mtry = m, # current candidate value of mtry
importance = "permutation" # compute permutation-based variable importance
)
# Store the current mtry value and its OOB prediction error
rf_results <- bind_rows(
rf_results,
tibble(
mtry = m,
oob_error = fit$prediction.error
)
)
}
rf_results |>
paged_table()Plot the tuning results
rf_results |>
ggplot(aes(x = mtry, y = oob_error)) +
geom_line() +
geom_point() +
labs(
title = "Random Forest Tuning Results",
x = "mtry",
y = "Out-of-bag prediction error"
)
Final random forest
best_mtry <- rf_results |>
slice_min(oob_error, n = 1) |>
pull(mtry)
final_boston_rf <- ranger(
medv ~ .,
data = dtrain,
num.trees = 500,
mtry = best_mtry,
importance = "permutation"
)
best_mtry[1] 10final_boston_rfRanger result
Call:
ranger(medv ~ ., data = dtrain, num.trees = 500, mtry = best_mtry, importance = "permutation")
Type: Regression
Number of trees: 500
Sample size: 392
Number of independent variables: 13
Mtry: 10
Target node size: 5
Variable importance mode: permutation
Splitrule: variance
OOB prediction error (MSE): 9.527965
R squared (OOB): 0.8858911 Test performance of the random forest
rf_test_pred <- predict(final_boston_rf, data = dtest)$predictions
sqrt(mean((y_test - rf_test_pred)^2))[1] 3.075023Random forest variable importance
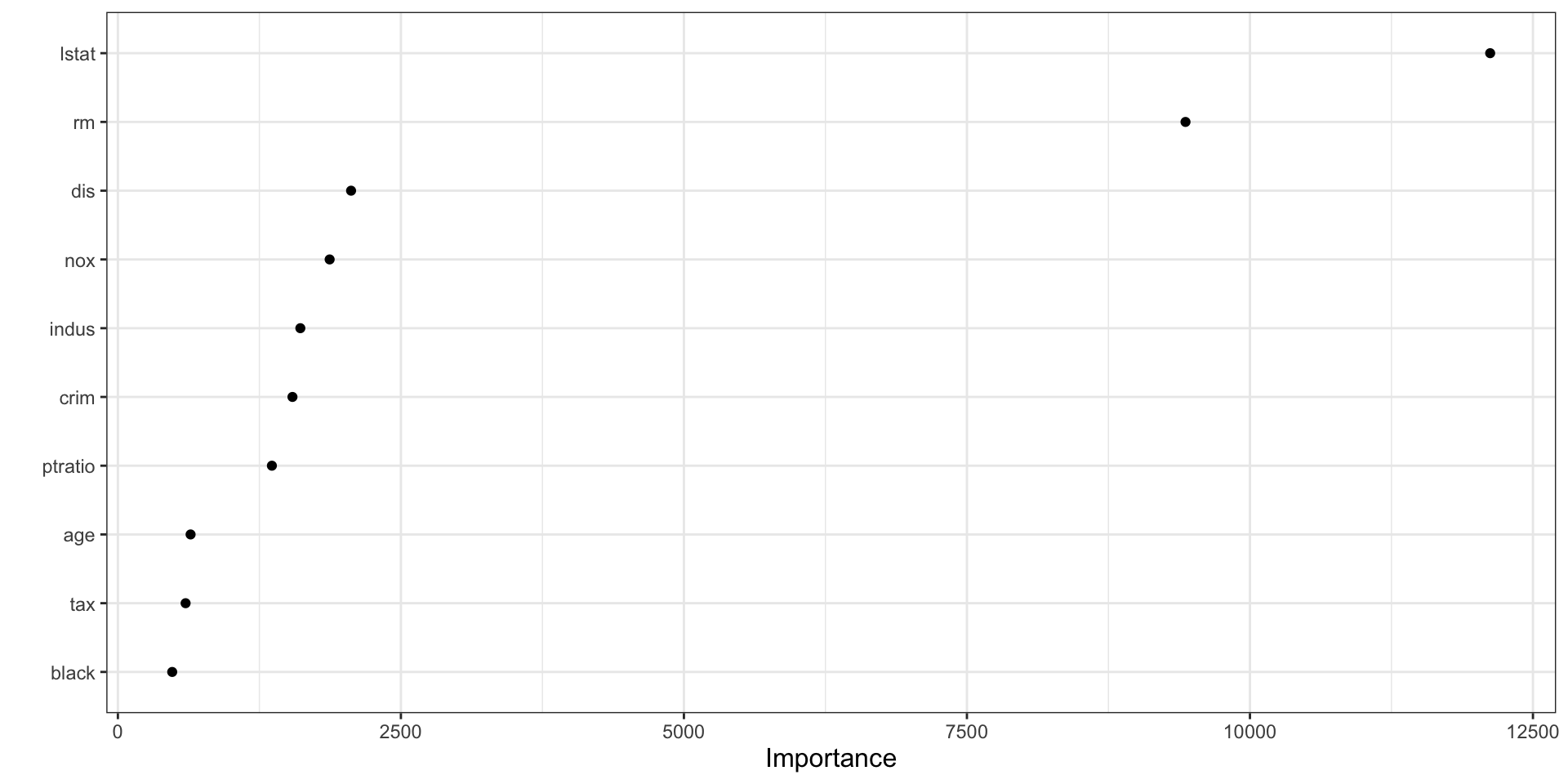
vip(final_boston_rf)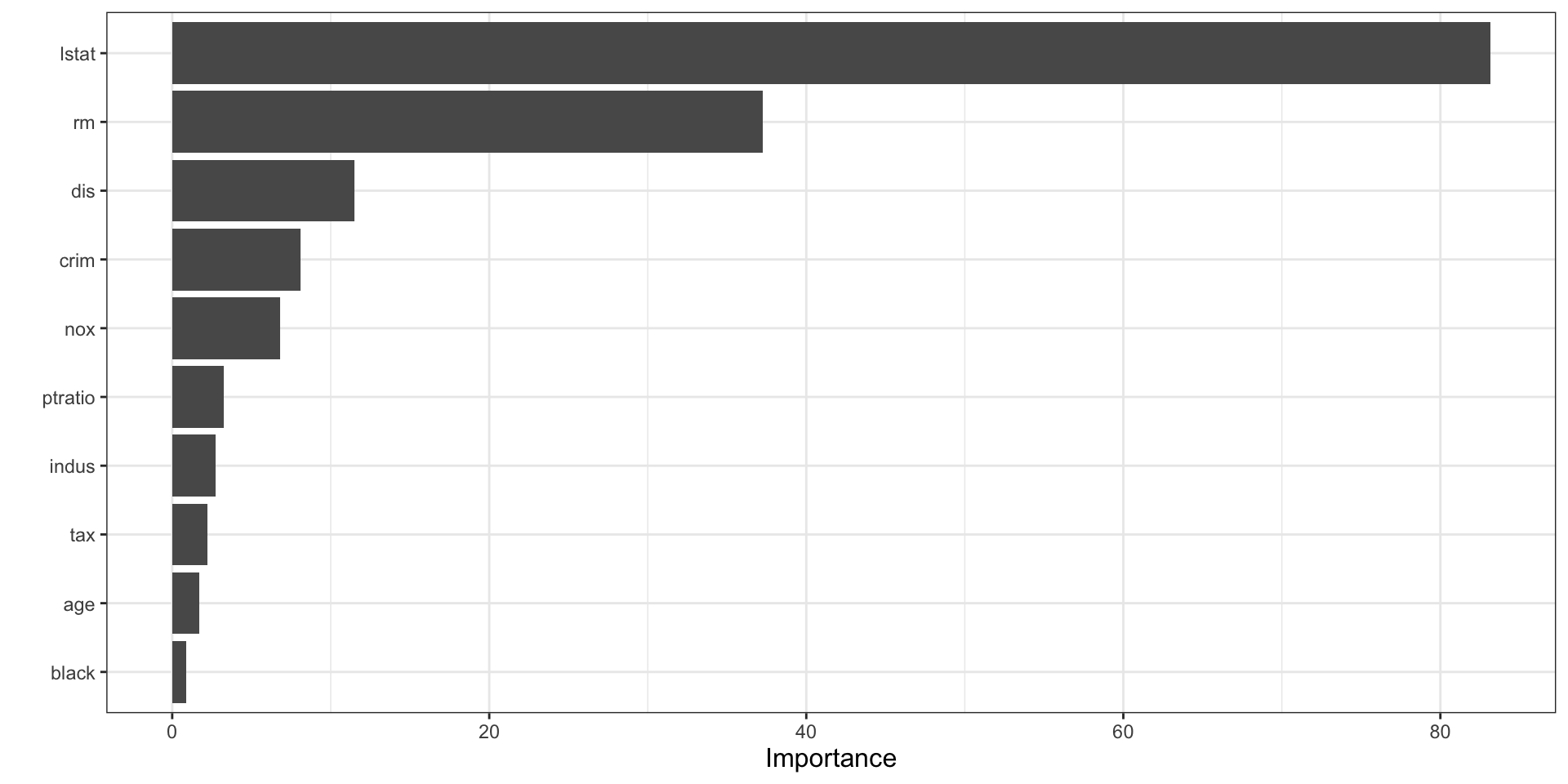
How to interpret vip() for a random forest
- The plot shows which predictors contributed most across the ensemble of trees.
- Higher importance means a variable was more useful in improving splits across the forest.
- In forests, importance is often more stable than in a single tree because it aggregates over many trees.
Partial dependence from the forest
partial(final_boston_rf, pred.var = "lstat", train = x_train) |>
autoplot() +
labs(title = "Partial Dependence of MEDV on LSTAT")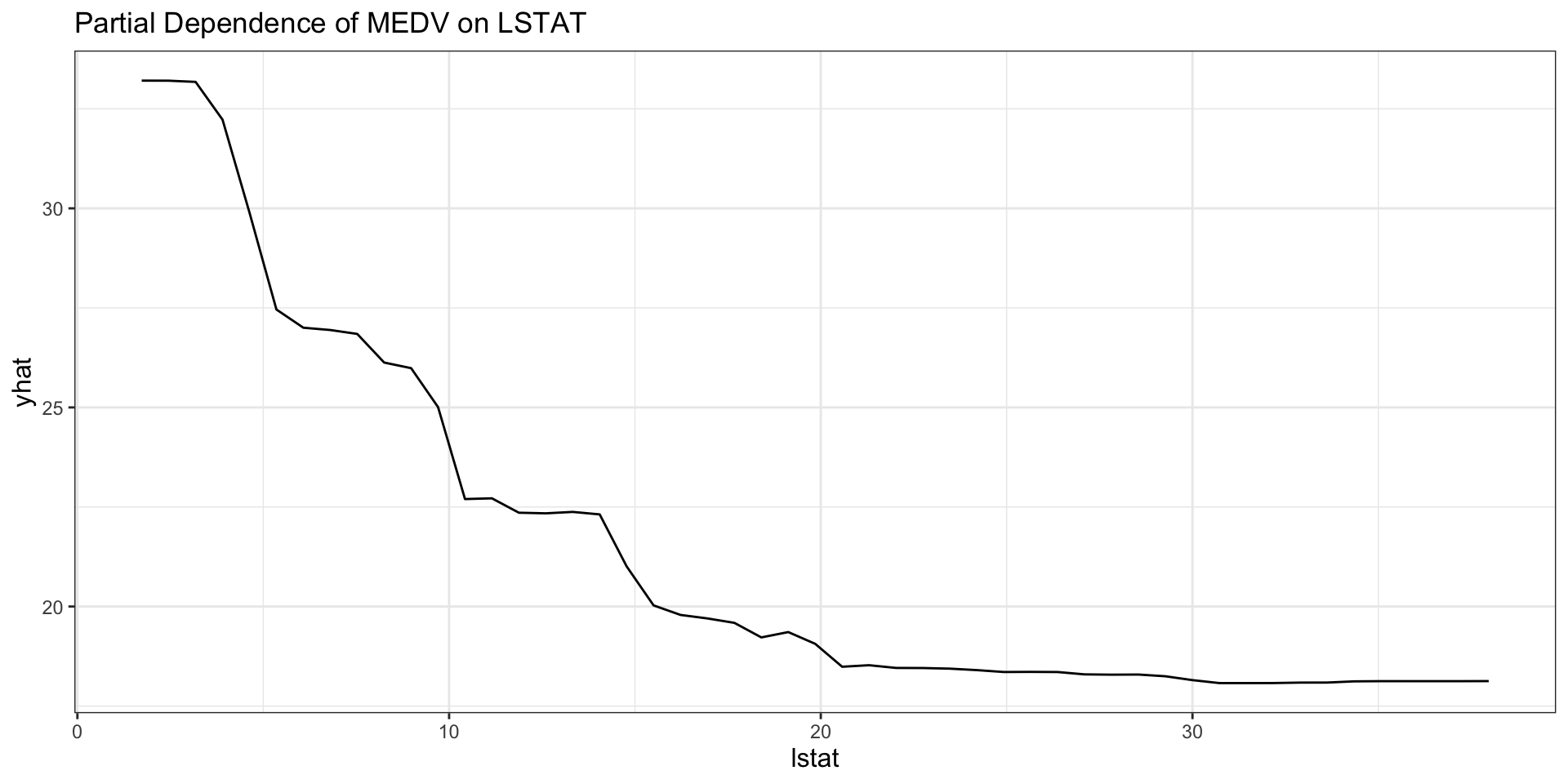
Interpreting the random-forest PDP
- The curve shows the model’s average predicted
medvaslstatchanges. - All other variables are averaged over their observed values.
- Nonlinearity in the curve suggests that the marginal relationship is not well described by a straight line.
- Flat regions suggest limited marginal predictive contribution in that range.
Gradient-Boosted Trees
Key XGBoost tuning arguments
nrounds: number of boosting iterationseta: learning ratemax_depth: depth of each treegamma: minimum improvement required for a splitsubsample: fraction of observations used for each boosting stepcolsample_bytree: fraction of predictors used for each treemin_child_weight: minimum total weight allowed in a child node
How to think about the main tuning parameters
- Smaller
etameans slower learning and usually requires largernrounds. - Larger
max_depthallows more complex interactions. - Larger
gammamakes splitting more conservative. - Smaller
subsampleandcolsample_bytreecan reduce overfitting by injecting randomness. nroundsandetausually need to be tuned together.
Prepare matrix input for XGBoost
x_train_mat <- as.matrix(x_train)
x_test_mat <- as.matrix(x_test)Fit an initial XGBoost model
library(xgboost)
set.seed(1937)
xgb_fit_initial <- xgboost(
data = x_train_mat,
label = y_train,
objective = "reg:squarederror",
nrounds = 100,
eta = 0.1,
max_depth = 3,
verbose = 0
)
xgb_fit_initialXGBoost model object
Call:
xgboost(objective = "reg:squarederror", nrounds = 100, max_depth = 3,
data = x_train_mat, label = y_train, eta = 0.1, verbose = 0)
Objective: reg:squarederror
Number of iterations: 100
Number of features: 13What does the initial model represent?
- It is a sum of many shallow trees, not one large tree.
- Each boosting round adds a small correction to the current prediction.
objective in xgboost()
objective = "reg:squarederror"is used for a regression problem, where the response variable is continuous.- It tells XGBoost to predict a numeric outcome and minimize squared prediction error.
objective = "binary:logistic"is used for a binary classification problem, where the response variable has two classes such as0and1.- It tells XGBoost to predict the probability that an observation belongs to class
1.
- It tells XGBoost to predict the probability that an observation belongs to class
Simple tuning grid
xgb_grid <- tidyr::crossing(
nrounds = seq(50, 250, by = 50),
eta = c(0.025, 0.05, 0.1),
max_depth = c(1, 2, 3, 4),
gamma = c(0, 1),
colsample_bytree = c(0.8, 1),
min_child_weight = 1,
subsample = c(0.8, 1)
)
xgb_grid |>
paged_table()tidyr::crossing() generates all possible combinations of the supplied values, which is exactly what you want for a manual grid search.
- Grid size: 5 × 3 × 4 × 2 × 2 × 1 × 2 = 480 combinations
min_child_weight = 1is held constant (scalar), while everything else is varied- Here,
nroundsandetaare treated independently — combinations likenrounds = 50, eta = 0.025(very slow learning, few rounds) will both be evaluated alongsidenrounds = 250, eta = 0.1- We can consider fixing
etaand using early stopping instead, which makesnroundsa ceiling rather than a tuning parameter
- We can consider fixing
Cross-validated tuning
xgb_results_lst <- vector("list", nrow(xgb_grid))
dMat_train <- xgb.DMatrix(data = x_train_mat, label = y_train)
for (i in seq_len(nrow(xgb_grid))) {
nrounds <- xgb_grid$nrounds[i]
eta <- xgb_grid$eta[i]
max_depth <- xgb_grid$max_depth[i]
gamma <- xgb_grid$gamma[i]
colsample_bytree <- xgb_grid$colsample_bytree[i]
min_child_weight <- xgb_grid$min_child_weight[i]
subsample <- xgb_grid$subsample[i]
cv_fit <- xgb.cv(
# data = x_train_mat,
# label = y_train,
# If an error occurs with the message including DMatrix, try:
data = dMat_train,
nrounds = nrounds,
nfold = 5,
objective = "reg:squarederror",
eta = eta,
max_depth = max_depth,
gamma = gamma,
colsample_bytree = colsample_bytree,
min_child_weight = min_child_weight,
subsample = subsample,
verbose = 0
)
xgb_results_lst[[i]] <- tibble(
nrounds = nrounds,
eta = eta,
max_depth = max_depth,
gamma = gamma,
colsample_bytree = colsample_bytree,
min_child_weight = min_child_weight,
subsample = subsample,
cv_rmse = min(cv_fit$evaluation_log$test_rmse_mean)
)
}
xgb_results <-
bind_rows(xgb_results_lst) |>
arrange(cv_rmse)
xgb_results |>
paged_table()Fit the final XGBoost model
# best_xgb <- xgb_results[order(xgb_results$cv_rmse), ][1, ]
best_xgb <- xgb_results |>
slice_min(cv_rmse, n = 1)
best_xgb |>
paged_table()xgb_fit_final <- xgboost(
data = x_train_mat,
label = y_train,
objective = "reg:squarederror",
nrounds = best_xgb$nrounds,
eta = best_xgb$eta,
max_depth = best_xgb$max_depth,
gamma = best_xgb$gamma,
colsample_bytree = best_xgb$colsample_bytree,
min_child_weight = best_xgb$min_child_weight,
subsample = best_xgb$subsample,
verbose = 0
)
xgb_fit_finalXGBoost model object
Call:
xgboost(objective = "reg:squarederror", nrounds = best_xgb$nrounds,
max_depth = best_xgb$max_depth, min_child_weight = best_xgb$min_child_weight,
subsample = best_xgb$subsample, colsample_bytree = best_xgb$colsample_bytree,
data = x_train_mat, label = y_train, eta = best_xgb$eta,
gamma = best_xgb$gamma, verbose = 0)
Objective: reg:squarederror
Number of iterations: 150
Number of features: 13Test performance of XGBoost
xgb_test_pred <- predict(xgb_fit_final, newdata = x_test_mat)
sqrt(mean((y_test - xgb_test_pred)^2))[1] 2.841075Variable Importance: A Free Byproduct of Ensembles
Both random forest and gradient boosting measure variable importance automatically
The key question: how much does each predictor contribute to reducing prediction error?
Random forest — permutation importance:
- For each variable, randomly shuffle its values and measure the drop in OOB accuracy
- A large drop → the variable was doing real work; a small drop → it barely mattered
Random forest — impurity importance:
- Tracks how much splits on a variable reduce impurity, summed across all trees
- In regression, impurity is usually based on reduction in outcome variance
- A larger total reduction means the variable was more useful for making cleaner splits
Gradient boosting — gain-based importance:
- Tracks how much each split on a variable reduced the loss function, summed across all trees
- Variables used in high-gain splits rank highly
Why This Matters
Unlike linear regression, tree ensembles can capture non-linear effects and interactions. Variable importance tells you which variables the model relies on most, even when the relationship between predictor and outcome is complex.
XGBoost variable importance
vip(xgb_fit_final) +
labs(title = "XGBoost Variable Importance")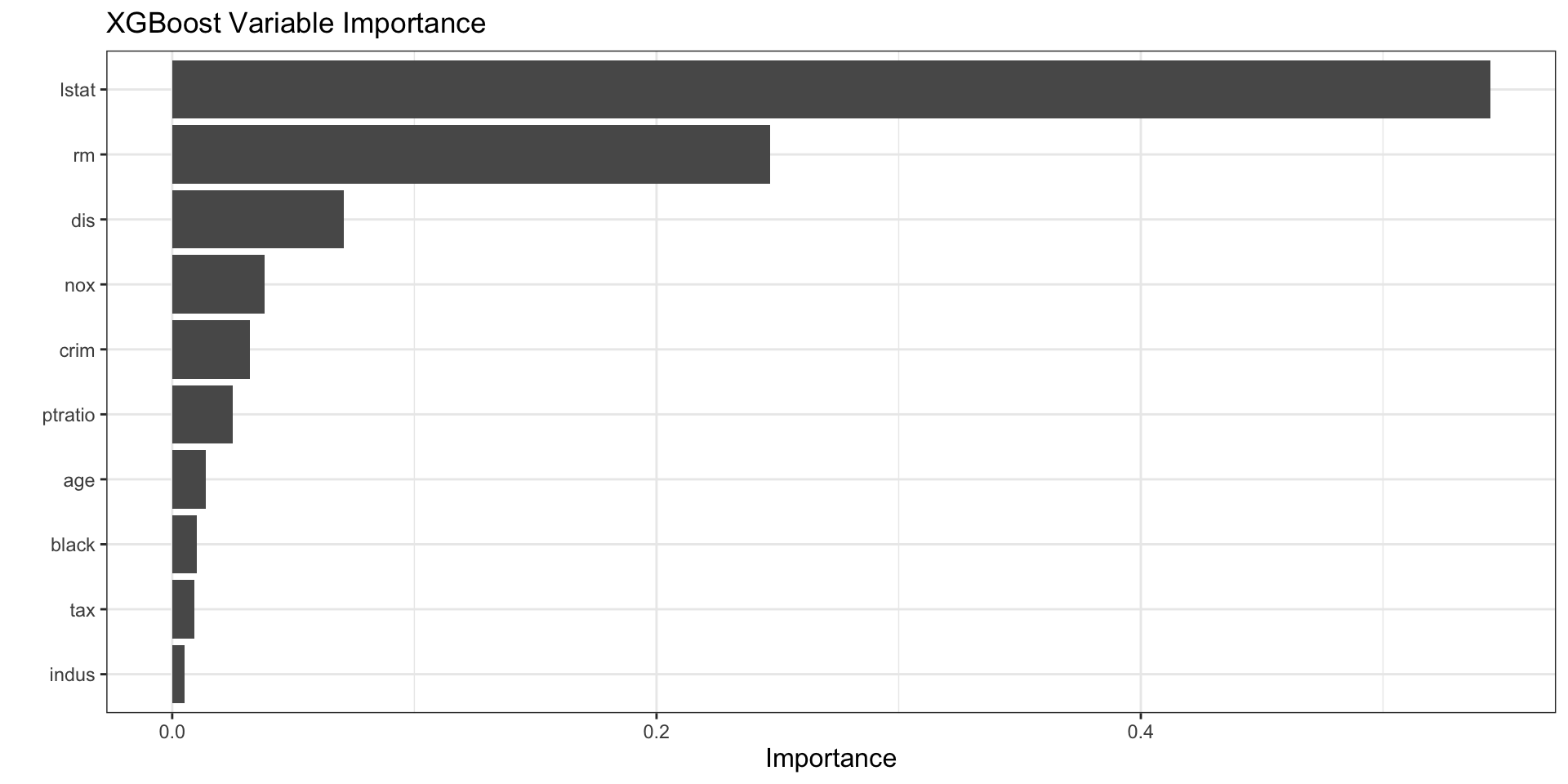
How to interpret vip() for XGBoost
- XGBoost importance is based on how much each variable contributes across many boosting rounds.
- Variables near the top are most useful for reducing the loss function.
- Importance does not reveal the direction of the effect.
- A variable can be important because it appears in many small improvements, not only because it appears in one dominant split.
Partial dependence for XGBoost
partial(
xgb_fit_final,
pred.var = "lstat",
train = x_train_mat,
plot = TRUE,
plot.engine = "ggplot2",
type = "regression"
) +
labs(title = "Partial Dependence of MEDV on LSTAT")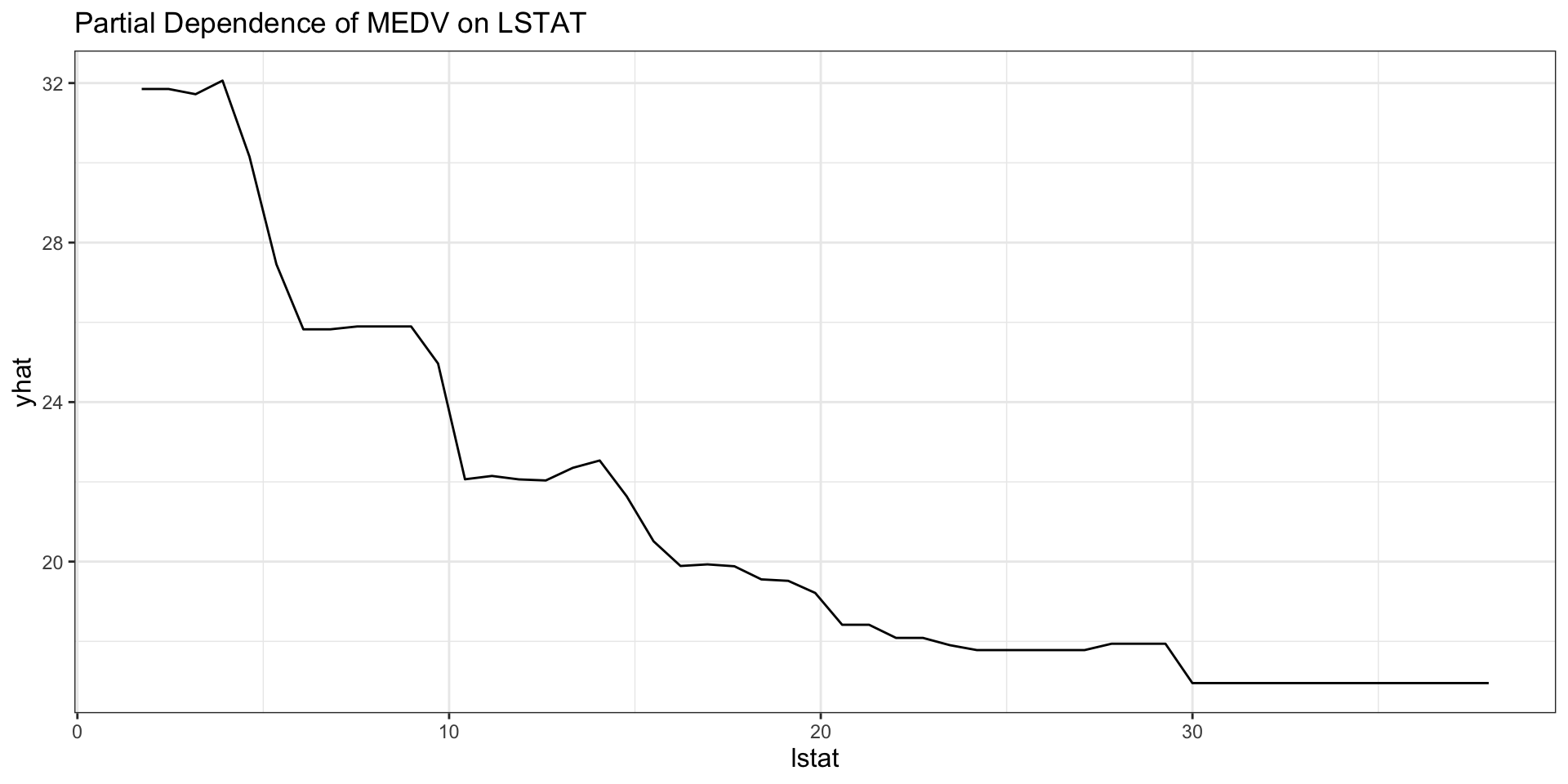
Interpreting the XGBoost PDP
- The PDP summarizes the average fitted relationship between
lstatand predicted housing values. - Because XGBoost is highly flexible, the curve can show richer nonlinearities than a single tree.
- Sharp changes in the curve can indicate thresholds learned by the boosted ensemble.
